Intragenomic Decarboxylation of 5-Carboxy-2′-deoxycytidine
2021-08-25
Ewelina Kamińska, M. Sc. Eva Korytiaková, M. Sc. Andreas Reichl, Dr. Markus Müller, Prof. Dr. Thomas Carell
Angew. Chem. Int. Ed., 2021, 60, 23207 –23211
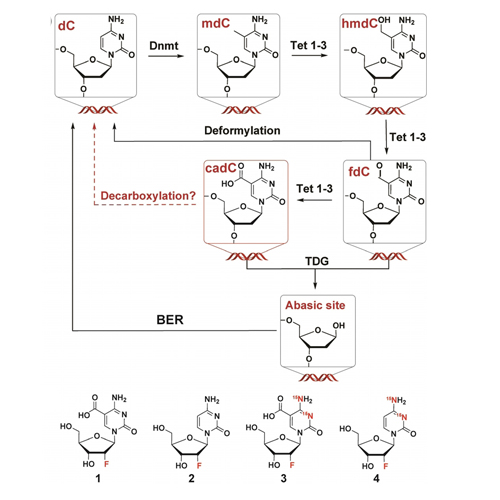
Cellular DNA is composed of four canonical nucleosides (dA, dC, dG and T), which form two Watson–Crick base pairs. In addition, 5-methylcytosine (mdC) may be present. The methylation of dC to mdC is known to regulate transcriptional activity. Next to these five nucleosides, the genome, particularly of stem cells, contains three additional dC derivatives, which are formed by stepwise oxidation of the methyl group of mdC with the help of Tet enzymes. These are 5-hydroxymethyl-dC (hmdC), 5-formyl-dC (fdC), and 5-carboxy-dC (cadC). It is believed that fdC and cadC are converted back into dC, which establishes an epigenetic control cycle that starts with methylation of dC to mdC, followed by oxidation and removal of fdC and cadC. While fdC was shown to undergo intragenomic deformylation to give dC directly, a similar decarboxylation of cadC was postulated but not yet observed on the genomic level. By using metabolic labelling, we show here that cadC decarboxylates in several cell types, which confirms that both fdC and cadC are nucleosides that are directly converted back to dC within the genome by C−C bond cleavage.